Optimized Breast Cancer Classification using Feature Selection and Outliers Detection
Abstract
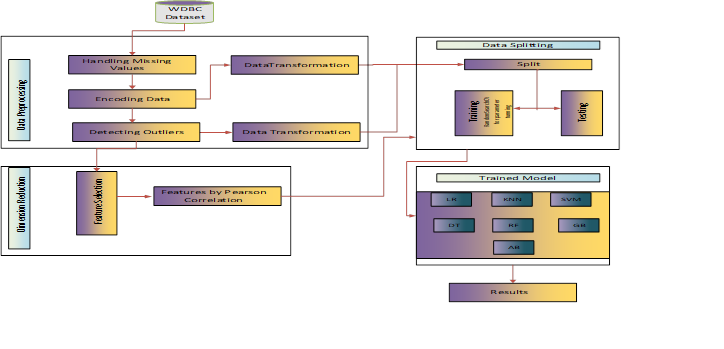
Breast cancer is the second most commonly diagnosed cancer in women throughout the world. It is on the rise, especially in developing countries, where the majority of cases are discovered late. Breast cancer develops when cancerous tumors form on the surface of the breast cells. The absence of accurate prognostic models to assist physicians recognize symptoms early makes it difficult to develop a treatment plan that would help patients live longer. However, machine learning techniques have recently been used to improve the accuracy and speed of breast cancer diagnosis. If the accuracy is flawless, the model will be more efficient, and the solution to breast cancer diagnosis will be better. Nevertheless, the primary difficulty for systems developed to detect breast cancer using machine-learning models is attaining the greatest classification accuracy and picking the most predictive feature useful for increasing accuracy. As a result, breast cancer prognosis remains a difficulty in today's society. This research seeks to address a flaw in an existing technique that is unable to enhance classification of continuous-valued data, particularly its accuracy and the selection of optimal features for breast cancer prediction. In order to address these issues, this study examines the impact of outliers and feature reduction on the Wisconsin Diagnostic Breast Cancer Dataset, which was tested using seven different machine learning algorithms. The results show that Logistic Regression, Random Forest, and Adaboost classifiers achieved the greatest accuracy of 99.12%, on removal of outliers from the dataset. Also, this filtered dataset with feature selection, on the other hand, has the greatest accuracy of 100% and 99.12% with Random Forest and Gradient boost classifiers, respectively. When compared to other state-of-the-art approaches, the two suggested strategies outperformed the unfiltered data in terms of accuracy. The suggested architecture might be a useful tool for radiologists to reduce the number of false negatives and positives. As a result, the efficiency of breast cancer diagnosis analysis will be increased.
Published
How to Cite
Issue
Section
Copyright (c) 2021 Journal of the Nigerian Society of Physical Sciences

This work is licensed under a Creative Commons Attribution 4.0 International License.






