Degradation of PET Nanoplastic Oligomers at the Novel PHL7 Target:Insights from Molecular Docking and Machine Learning
Keywords:
Polyethylene terephthalate, Nanoplastic, Polyester Hydrolase Leipzig 7, binding affinity, Artificial Neutral NetworkAbstract
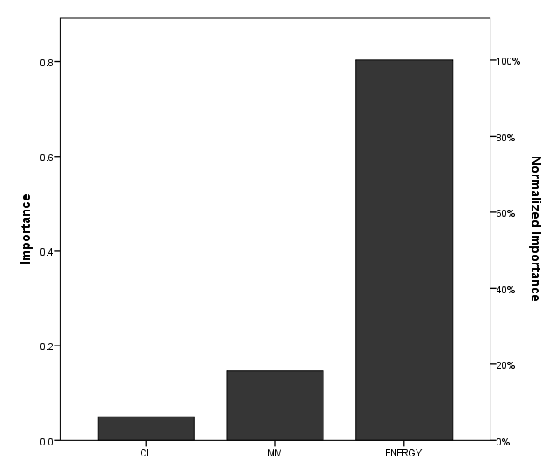
The versatility of Polyethylene terephthalate (PET) as a material with numerous applications in the food industry and its recalcitrance to chemical and microbial degradation has recently made it an environmental nuisance. In this study, we applied computational methods to ascertain the dependence of PET nanoplastic (NP) degradation on the chain length of the oligomer. The binding affinities of the NPs on the novel enzyme Polyester Hydrolase Leipzig 7 (PHL7) were used to relate their ease of degradation at the enzyme active site. The results revealed that the binding affinity of PET NPs at the enzyme target decreased from -5.2 kcal/mol to -0.8 kcal/mol, with an increase in PET chain length from 2.18 nm to 5.45 nm (2-5 PET chains). The binding affinities became positive at chain lengths 6.54 nm (6 PET chains) and above. These findings indicated that PET NP degradation at this enzyme’s active site is most efficient as chain length decreases from 5-2 units and is not likely to occur at longer PET chains. A feedforward Artificial Neutral Network (ANN) analysis predicted that the energy of the PET NPs is a very important factor in its degradation.
Published
How to Cite
Issue
Section
Copyright (c) 2023 C. E. Duru, C. E. Enyoh, I. A. Duru, M. C. Enedoh

This work is licensed under a Creative Commons Attribution 4.0 International License.
How to Cite
Similar Articles
- Emmanuel C. Ukekwe, Adaora A. Obayi, Akpa Johnson, Daniel A. Musa, Jonathan C. Agbo, Optimizing data and voice service delivery for mobile phones based on clients' demand and location using affinity propagation machine learning , Journal of the Nigerian Society of Physical Sciences: Volume 7, Issue 2, May 2025
- Hamza Abubakar, Abdu Sagir Masanawa, Surajo Yusuf, G. I. Boaku, Optimal representation to High Order Random Boolean kSatisability via Election Algorithm as Heuristic Search Approach in Hopeld Neural Networks , Journal of the Nigerian Society of Physical Sciences: Volume 3, Issue 3, August 2021
- L. G. Salaudeen, D. GABI, M. Garba, H. U. Suru, Deep convolutional neural network based synthetic minority over sampling technique: a forfending model for fraudulent credit card transactions in financial institution , Journal of the Nigerian Society of Physical Sciences: Volume 6, Issue 2, May 2024
- Nassima Bou-ydia, Ben-issa El miraoui, Latifa Laallam, Ahmed Jouaiti, Interaction of hydroxyapatite and chitosan with gentamicin and their antimicrobial activities: DFT and molecular docking approach , Journal of the Nigerian Society of Physical Sciences: Volume 7, Issue 3, August 2025
- Amos Orenyi Bajeh, Mary Olayinka Olaoye, Fatima Enehezei Usman-Hamza, Ikeola Suhurat Olatinwo, Peter ogirima Sadiku, Abdulkadir Bolakale Sakariyah, An adaptive neuro-fuzzy inference system for multinomial malware classification , Journal of the Nigerian Society of Physical Sciences: Volume 7, Issue 1, February 2025
- R. El chaal, M. O. Aboutafail, Statistical Modelling by Topological Maps of Kohonen for Classification of the Physicochemical Quality of Surface Waters of the Inaouen Watershed Under Matlab , Journal of the Nigerian Society of Physical Sciences: Volume 4, Issue 2, May 2022
- Chidi Duru, Ijeoma Duru, Chiagoziem Chidiebere, Virtual Screening of Selected Natural Products as Human Tyrosinase-Related Protein 1 Blockers , Journal of the Nigerian Society of Physical Sciences: Volume 3, Issue 3, August 2021
- U. B. Amadi, M. Ogwuegbu, C. K. Enenebeaku, G. onyedika, Design, synthesis and studies on thermodynamic and biological activities of lanthanide III complexes of hydrazinecarbothioamide ligands , Journal of the Nigerian Society of Physical Sciences: Volume 7, Issue 2, May 2025
- S. I. Ele, U. R. Alo, H. F. Nweke, A. H. Okemiri, E. O. Uche-Nwachi, Deep convolutional neural network (DCNN)-based model for pneumonia detection using chest x-ray images , Journal of the Nigerian Society of Physical Sciences: Volume 7, Issue 2, May 2025
- Wilson Nwankwo, Kingsley Ukhurebor, Investigating the Performance of Point to Multipoint Microwave Connectivity across Undulating Landscape during Rainfall , Journal of the Nigerian Society of Physical Sciences: Volume 1, Issue 3, August 2019
You may also start an advanced similarity search for this article.
Most read articles by the same author(s)
- Chidi Duru, Ijeoma Duru, Chiagoziem Chidiebere, Virtual Screening of Selected Natural Products as Human Tyrosinase-Related Protein 1 Blockers , Journal of the Nigerian Society of Physical Sciences: Volume 3, Issue 3, August 2021






